How to Get Started Analyzing COVID-19 Data
Mar 19, 2020 13:49 · 1417 words · 7 minute read
 Photo by National Cancer Institute on Unsplash
Photo by National Cancer Institute on Unsplash
Earlier this month, Kaggle released a new dataset challenge: the COVID-19 Open Research Dataset Challenge. This challenge is a call to action to AI experts to develop text processing tools to help medical professionals find answers to high priority questions.
To that end, and in partnership with AI2, CZI, MSR, Georgetown, NIH & The White House, Kaggle assembled a dataset “of over 29,000 scholarly articles, including over 13,000 with full text, about COVID-19, SARS-CoV-2, and related coronaviruses.”
I believe that a lot of data scientists are looking for opportunities to help combat the COVID-19 pandemic and this is a great place to start. Kaggle provides the data and even a set of tasks for you to tackle. For example, one task is:
What do we know about COVID-19 risk factors? What have we learned from epidemiological studies?
To help others get started with this dataset, I thought I would provide a fairly simple approach I took to analyzing the data. It’s important to note that this approach does not aim to be a comprehensive review of the data or the tasks. Instead, just one idea being walked through to help you hopefully get started with your idea.
Meta Data
I want to focus my efforts on the CORD-19-research-challenge/2020–03–13/all_sources_metadata_2020–03–13.csv file provided. You can read that file into a Pandas data frame with the following code:
meta = pd.read_csv("/kaggle/input/CORD-19-research-challenge/2020-03-13/all_sources_metadata_2020-03-13.csv")
Here are the first 5 rows of the metadata file:
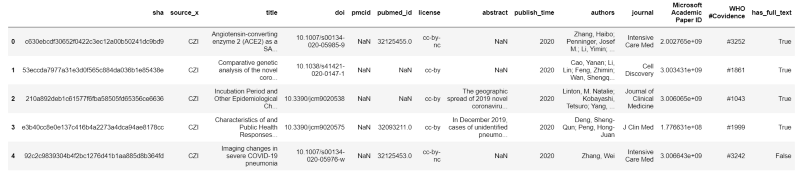
We will look exclusively at the abstract column. There are 29,500 rows of data in this file and 2,947 have missing abstracts (about 10 percent). Here are some basic statistics for the length (number of characters) of abstracts:
count 26553.000000
mean 1462.129176
std 1063.536406
min 17.000000
25% 1087.000000
50% 1438.000000
75% 1772.000000
max 122392.000000
Abstract Vectors
My idea is a simple one:
- Convert the abstracts into vectors
- Calculate cosine similarities to find abstracts similar to other abstracts
The goal is that if a medical researcher were to find a valuable paper he or she could take that paper’s abstract and quickly find other abstracts that are similar.
To calculate our abstract vectors we will leverage scispaCy. scispaCy is a “Python package containing spaCy models for processing biomedical, scientific or clinical text.” I decided to use this package created by allenai because it was trained specifically for biomedical text and is very easy to use. We can calculate vectors for all of our abstracts with the following code:
nlp = spacy.load(“en_core_sci_sm”)
vector_dict = {}
for sha, abstract in tqdm(meta[[“sha”,“abstract”]].values):
if isinstance(abstract, str):
vector_dict[sha] = nlp(abstract).vector
This code leverages a pre-trained word vector model that converts every word in the abstract to a vector representation which hopefully is a good representation of that word. A good representation would be close in vector space to other words which are similar (at a very high level). All the word vectors are then averaged to represent the abstract as a single vector.
I use the sha values as lookup keys so we can easily connect the vectors to papers. I also used the small version of the model in order to quickly process all of the abstracts. It took about 20 minutes to run the above code on a Kaggle Kernel.
Cosine Similarity
Now that we have all of the abstracts represented as vectors, it is easy to use sklearn to calculate all the pairwise cosine similarities. Cosine similarity is a method to compare the similarity of two vectors.
values = list(vector_dict.values())
cosine_sim_matrix = cosine_similarity(values, values)
Each value of our cosine similarity matrix is the similarity score between two abstracts (the abstract represented by the row, and the abstract represented by the column). The higher the value, the more similar the abstracts (assuming our vectors are good). Thus, we just need to select a row (abstract) and sort it by the indexes with the largest values (ignoring its own index since it will always be most similar to itself). Here’s the code:
n_sim_articles = 5
input_sha = “e3b40cc8e0e137c416b4a2273a4dca94ae8178cc”
keys = list(vector_dict.keys())
sha_index = keys.index(input_sha)
sim_indexes = np.argsort(cosine_sim_matrix[sha_index])[::-1][1:n_sim_articles+1]
sim_shas = [keys[i] for i in sim_indexes]
meta_info = meta[meta.sha.isin(sim_shas)]
You just need to provide the number of similar articles you want and the input sha for your query abstract to get results. Let’s consider the following example abstract as a query:
In December 2019, cases of unidentified pneumonia with a history of exposure in the Huanan Seafood Market were reported in Wuhan, Hubei Province. A novel coronavirus, SARS-CoV-2, was identified to be accountable for this disease. Human-to-human transmission is confirmed, and this disease (named COVID-19 by World Health Organization (WHO)) spread rapidly around the country and the world. As of 18 February 2020, the number of confirmed cases had reached 75,199 with 2009 fatalities. The COVID-19 resulted in a much lower case-fatality rate (about 2.67%) among the confirmed cases, compared with Severe Acute Respiratory Syndrome (SARS) and Middle East Respiratory Syndrome (MERS). Among the symptom composition of the 45 fatality cases collected from the released official reports, the top four are fever, cough, short of breath, and chest tightness/pain. The major comorbidities of the fatality cases include hypertension, diabetes, coronary heart disease, cerebral infarction, and chronic bronchitis. The source of the virus and the pathogenesis of this disease are still unconfirmed. No specific therapeutic drug has been found. The Chinese Government has initiated a level-1 public health response to prevent the spread of the disease. Meanwhile, it is also crucial to speed up the development of vaccines and drugs for treatment, which will enable us to defeat COVID-19 as soon as possible.
According to our algorithm, the most similar abstract would be:
BACKGROUND: Until 2008, human rabies had never been reported in French Guiana. On 28 May 2008, the French National Reference Center for Rabies (Institut Pasteur, Paris) confirmed the rabies diagnosis, based on hemi-nested polymerase chain reaction on skin biopsy and saliva specimens from a Guianan, who had never travelled overseas and died in Cayenne after presenting clinically typical meningoencephalitis. METHODOLOGY/PRINCIPAL FINDINGS: Molecular typing of the virus identified a Lyssavirus (Rabies virus species), closely related to those circulating in hematophagous bats (mainly Desmodus rotundus) in Latin America. A multidisciplinary Crisis Unit was activated. Its objectives were to implement an epidemiological investigation and a veterinary survey, to provide control measures and establish a communications program. The origin of the contamination was not formally established, but was probably linked to a bat bite based on the virus type isolated. After confirming exposure of 90 persons, they were vaccinated against rabies: 42 from the case’s entourage and 48 healthcare workers. To handle that emergence and the local population’s increased demand to be vaccinated, a specific communications program was established using several media: television, newspaper, radio. CONCLUSION/SIGNIFICANCE: This episode, occurring in the context of a Department far from continental France, strongly affected the local population, healthcare workers and authorities, and the management team faced intense pressure. This observation confirms that the risk of contracting rabies in French Guiana is real, with consequences for population educational program, control measures, medical diagnosis and post-exposure prophylaxis.
If you look at the top 5 most similar abstracts, it seemed to connect the query article to articles discussing rabies. In my opinion, this doesn’t seem to be a great result. That being said, I am not a medical researcher so I might be missing something. But let’s assume this first pass is lacking in value.
Next Steps
I believe the idea represented above is sound. My guess is that the word vectors are not especially good for this body of research. As a next step, I would look at training word vectors on the text of all of the papers supplied. That should help the vectors be more tuned to this area of research.
There is also a lot of text processing that could be done to potentially improve the process. For example, removing stop words might help reduce the noise in some abstracts.
Vinit Jain, on LinkedIn, also suggested trying out Biobert or BioSentvec which I find to also be good ideas. Especially the idea of converting the abstracts to vectors in a smarter way than just averaging word vectors.
While my first attempt at providing value for the COVID-19 dataset was lackluster, I hope this article accomplished its true goal of helping you get started with analyzing the data.
You can find my Kaggle Kernel with all of the code discussed here.
Interested in learning more about Python data analysis and visualization? Check out my course.